Certain restriction enzymes break the double stranded DNA (dsDNA) at the centre of their recognition sites to produce blunt ended fragments that are based paired to their ends and thus have no tendency to stick together. These DNA fragments are referred to as blunt ends. For example the restriction enzyme HindII (isolated from E. coli) recognizes the specific sequence GTC/GAC (C represents the DNA base cytosine, G guanine, A adenine and T thymine). In contrast, the enzyme EcoRI makes staggered cuts that produce single stranded DNA (ssDNA) tails at the site of each cut. The recognition sequence of EcoRI (isolated from E. coli) is G/AATTC. Complimentary single-stranded tails (produced by the same cutting enzyme) are able to associate by base pairing of the DNA (hydrogen bonding occurs between adenine-thymine and guanine-cytosine) and are thus called sticky ends.


From left to right:
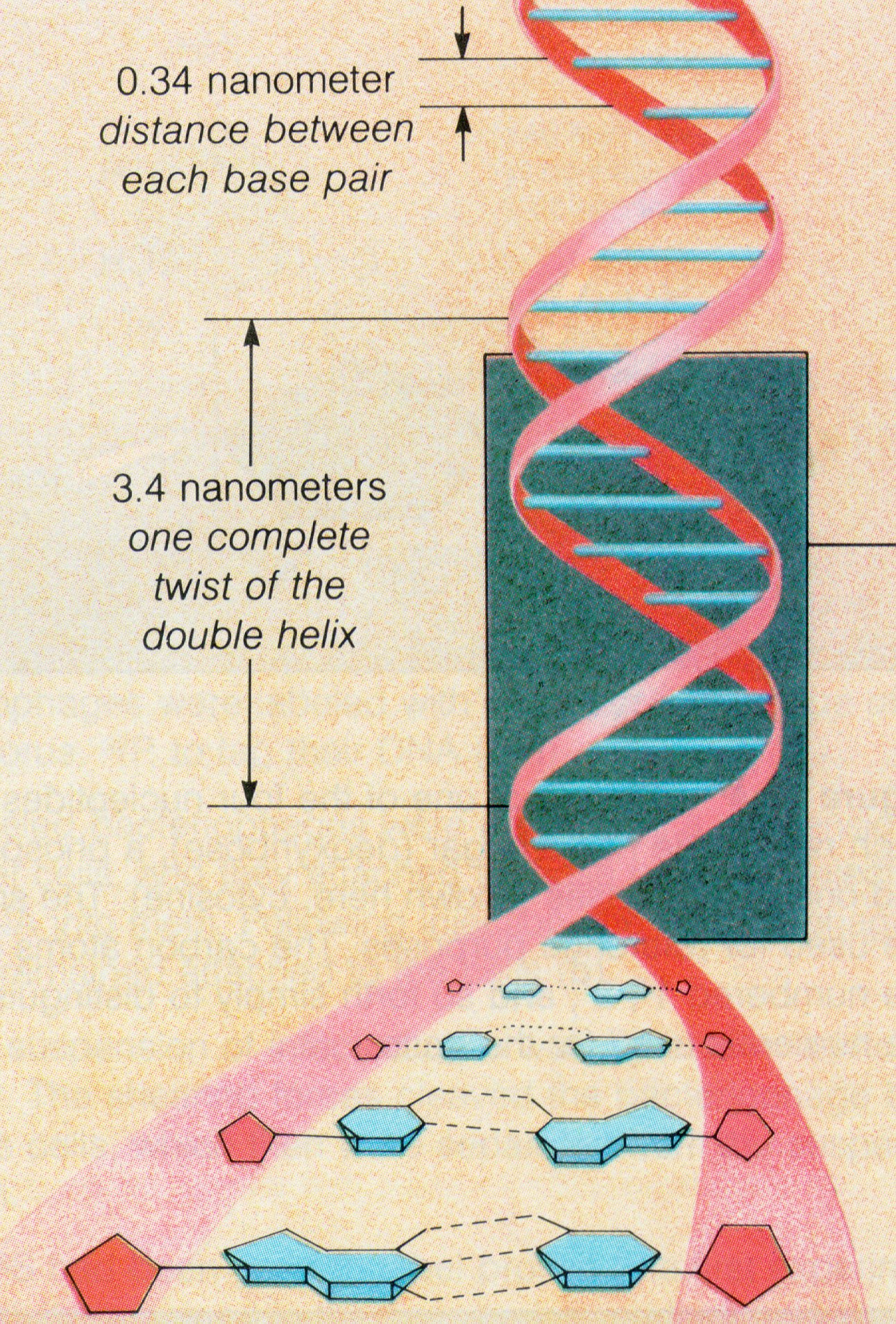
* The double stranded nature of the DNA helix
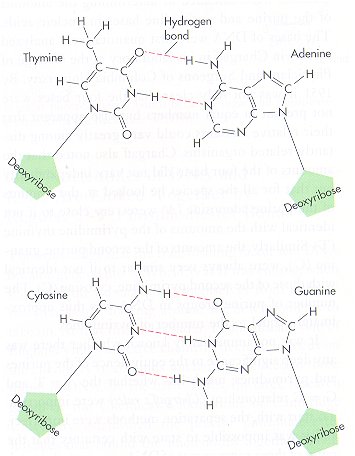
* The base pairing between adenine-thymine and guanine-cytosine
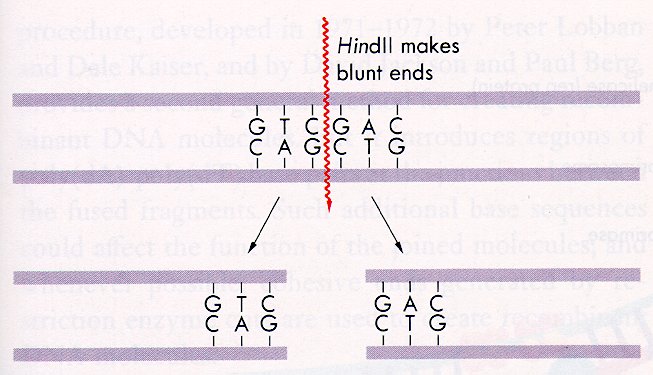
A Schematic of the blunt end incision made by HindII
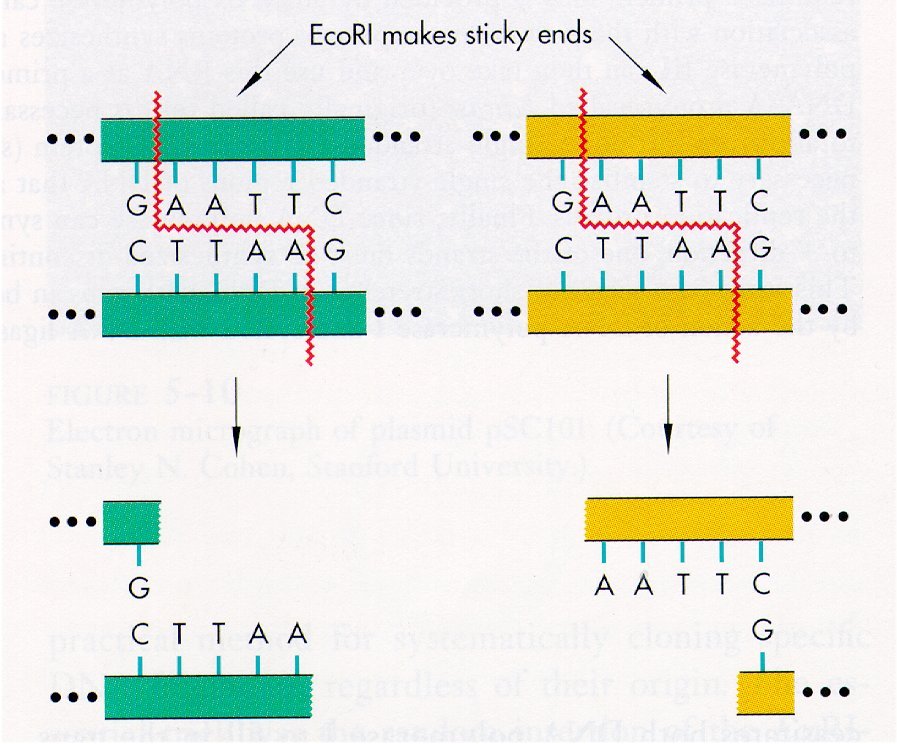
The sticky ends produced by an EcoRI incision and the subsequant ligation of the complimentary strands